We are exploring the use of cell mass and mass distribution measurements via quantitative phase imaging (QPI) to develop new approaches to study and treat human disease, including cancer. Our work includes collaborations with researchers at the University of Utah, Huntsman Cancer Institute, and other institutions. Code from these and other projects is available on GitHub.
Functional precision medicine
A critical problem in cancer care is how to match patients to the correct therapy. Despite advances in ‘omics’ approaches, the molecular profiling of tumors has not efficiently translated in effectively personalized therapies for the majority of cancer patients. An alternative approach, called functional precision medicine, directly tests the response of patient tumor cells when treated with a panel of candidate therapies to inform oncologists and match patients to the right therapy. However, current approaches to measure the response of tumor cells to chemotherapy require 1-8 months to obtain results, making these impractical for patient care.
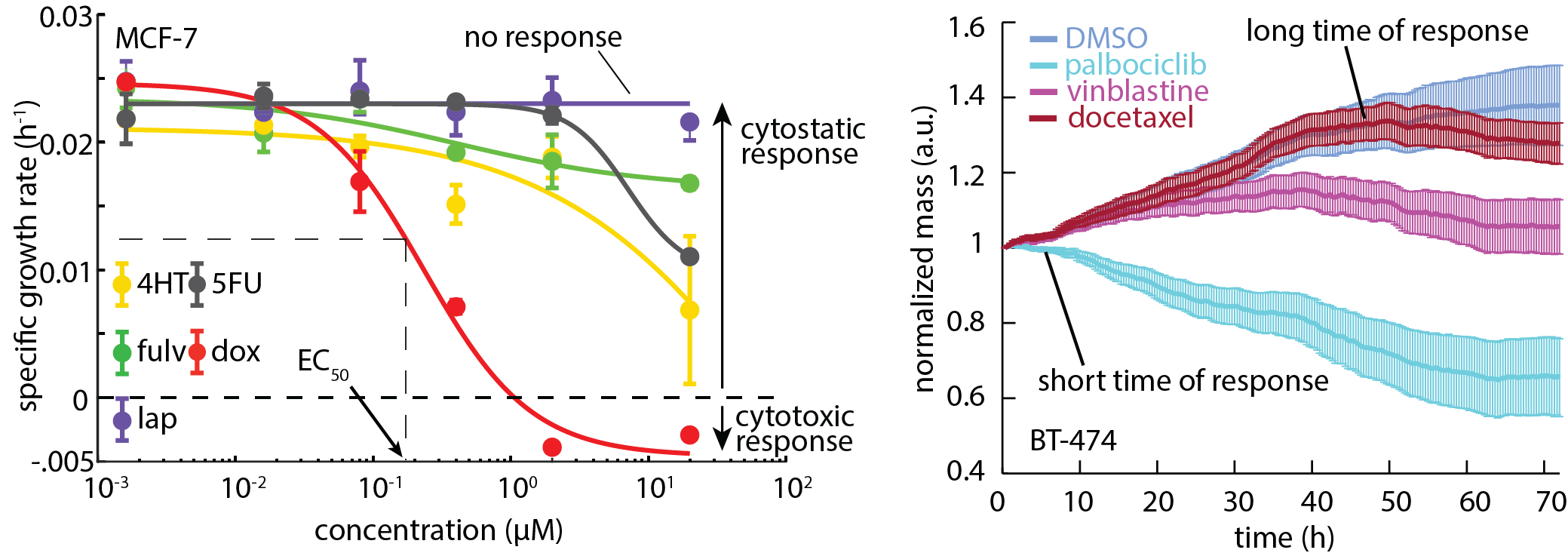
The Zangle lab, in collaboration with labs at the Huntsman Cancer Institute (HCI), has developed instruments and analytical methods based on QPI to predict cancer cell response to therapy within days of sample collection in terms of both population response and single-cell heterogeneity. The core principle is to use QPI to measure the rate of change of cell mass over time. We can then score potential therapies based on how much they slow or halt this growth. This allows QPI to rapidly measure the response of individual cells to therapy, without requiring labels, and with relatively small numbers of cells, therefore reducing the need for costly and time-consuming cell expansion, reducing overall assay time and cost. We have shown that this approach can be used to simultaneously measure multiple parameters that define cancer cell response to therapy, an approach we call Multimodal QPI. These parameters, including standard response parameters such as EC50 and depth of response, as well as new parameters such as time and heterogeneity of response, give a richer picture of cancer cell response to therapy which we are working to translate into a clinical impact on patient outcomes.
Key papers:
Edward R. Polanco, Tarek E. Moustafa, Andrew Butterfield, Sandra D. Scherer, Emilio Cortes-Sanchez, Tyler Bodily, Benjamin T. Spike, Bryan E. Welm, Philip S. Bernard, and Thomas A. Zangle, “Multiparametric quantitative phase imaging for real-time, single cell, drug screening in breast cancer,” bioRxiv 2021.11.26.467625. doi: https://doi.org/10.1101/2021.11.26.467625.
Devin Lange, Edward R. Polanco, Robert Judson-Torres, Thomas A. Zangle, and Alexander Lex, “Loon: Using exemplars to visualize large-scale microscopy data,” IEEE Transactions on Visualization and Computer Graphics (VIS) 2021. doi: 10.1109/TVCG.2021.3114766. (link) (pdf)
Thomas A. Zangle* and Michael A. Teitell,* “Live cell mass profiling: An emerging approach in quantitative biophysics,” Nature Methods 2014, 11. 1221-1228. *co-corresponding (link) (pdf)
Fabrication of refractive index-matched microdevices
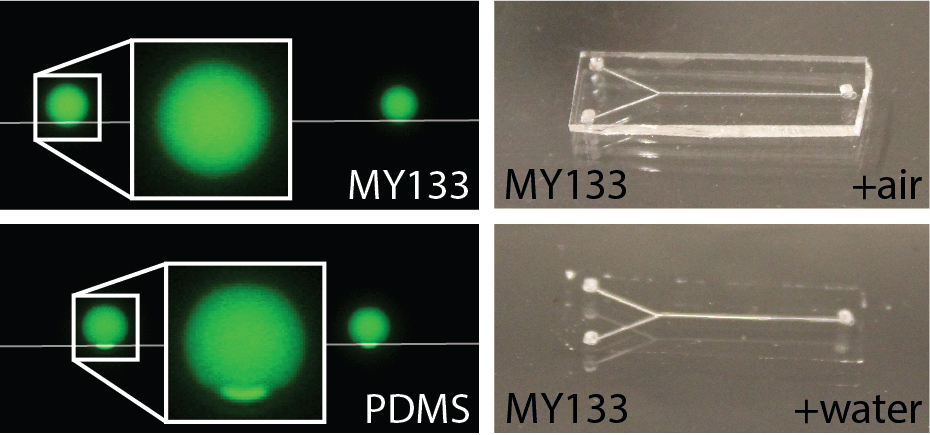
Conventional materials used to fabricate microfluidic devices (e.g. PDMS) typically have a high refractive index relative to water. This can lead to artifacts at device edges during imaging. The Zangle lab has developed fabrication approaches to create refractive-index matched microfluidic devices that can confine and manipulate cells without interfering with imaging. We have found that refractive index matching, while critical for effective QPI, is also useful for reducing artifacts in other imaging modalities, including fluorescence, where cells to be imaged are near device edges. Fluorination is typically used to reduce material polarizability and, therefore, refractive index. However, this comes at the cost of reduced material adhesion. The Zangle lab has therefore demonstrated a range of approaches to mitigate this impact and successfully bond refractive-index matched microfluidic devices. We are applying these methods to confine cells within microwells for high throughput assays.
Key papers:
Edward R. Polanco, Justin Griffin, and Thomas A. Zangle, “Fabrication and bonding of refractive index matched microfluidics for precise measurements of cell mass,” Polymers 2021, 13. 496. (pdf)
Edward R. Polanco, Nicholas Western, and Thomas A. Zangle, “Fabrication of refractive-index-matched devices for biomedical microfluidics,” Journal of Visualized Experiments (JoVE) 2018, 139. e58296. (link) (pdf) (video)
Diane N.H. Kim, Kevin T. Kim, Carolyn Kim, Michael A. Teitell, and Thomas A. Zangle, “Soft lithography fabrication of index-matched microfluidic devices for reducing artifacts in fluorescence and quantitative phase imaging,” Microfluidics and Nanofluidics 2018, 22. 1-11. (link)
Intracellular dynamics
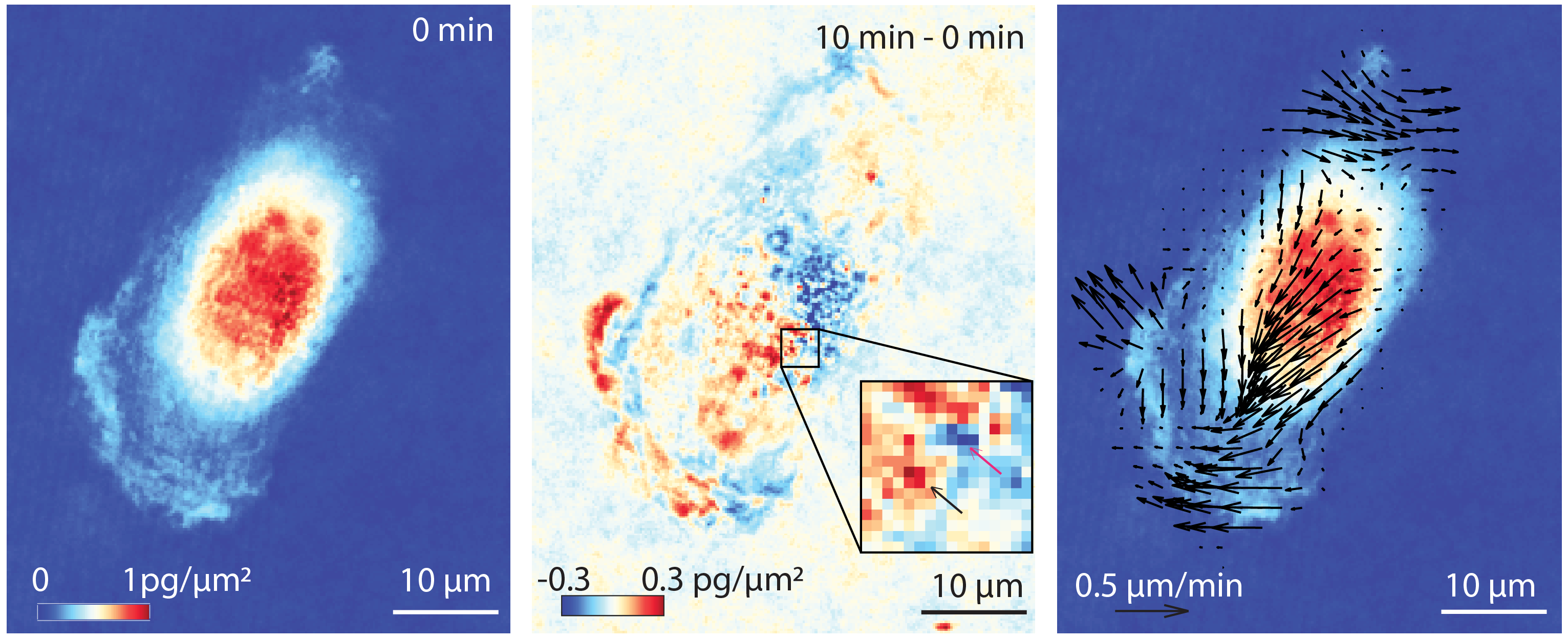
Most previous QPI studies measure total cell mass, ignoring the rich data on the motion of mass within the cell that is also acquired with this imaging approach. We are developing methods that leverage this data to develop a clearer picture of the dynamics of mass transport and growth processes within cells. We have used these data as the basis for a new, label-free approach to measure cell viscoelasticity. We have also developed methods to quantify the rate of neuronal body and neurite biomass accumulation. Finally, we have also developed a new method which we call quantitative phase velocimetry (QPV) to track the velocity of mass transport within cells. This approach enables tracing the path of individual parcels of mass as they move through the cell, and offers a potential alternative or complement to traditional micro rheology, in which individual particles or organelles are tracked within cells to measure intracellular properties. Ongoing work in this area will assess the impact of specific inhibitors on intracellular transport as a new modality for measuring cell drug responses.
Key papers:
Soorya Pradeep and Thomas A. Zangle, “LVING Reveals the Intracellular Structure of Cell Growth,” Scientific Reports, 2024, 14:8544. doi: 10.1038/s41598-024-58992-x. (link)
Soorya Pradeep and Thomas A. Zangle, “Quantitative phase velocimetry for label-free measurement of intracellular mass transport velocity,” Scientific Reports, 2022, 12. 6074. (link).
Soorya Pradeep, Tasmia Tasnim, Huanan Zhang, and Thomas A. Zangle, “Simultaneous measurement of neurite and neural body mass accumulation via quantitative phase imaging,” Analyst 2021, 146. 1361-1368. (link)
Thang L. Nguyen, Edward R. Polanco, Alexander N. Patananan, Thomas A. Zangle*, and Michael A. Teitell*, “Cell viscoelasticity is linked to fluctuations in cell biomass distributions,” Scientific Reports 2020, 10. 7403. *co-corresponding (link)
